
ここからコンテンツです。

Molecular modeling of novel potent agents for treating Alzheimer’s disease
Novel curcumin derivatives proposed for inhibiting the production of amyloid-beta peptides, which are the main causal agents of Alzheimer’s disease. By Noriyuki Kurita
Noriyuki Kurita and his colleagues in cooperation with the Ukraine National Academy of Sciences proposed novel agents for inhibiting the production of amyloid-beta (Aβ) peptides, which are involved in the pathogenesis of Alzheimer’s disease. Using state-of-the-art molecular simulations, interactions between amyloid precursor protein (APP) and curcumin derivatives were investigated, to elucidate the specific derivative that binds strongly to APP and inhibits the pathogenic Aβ production. The results contribute to developing new medicines that suppress Aβ peptide production.
Alzheimer's disease (AD), a severe form of dementia among aged individuals, is caused by accumulation of amyloid-beta (Aβ) peptides in the brain. Numerous types of agents have been developed to suppress the production of Aβ, by inhibiting the secretase-mediated cleavage of amyloid precursor protein (APP) into Aβ peptides.
However, because the secretases also play important roles in the production of vital proteins for the human body, inhibitors of the secretases have increased risk for side effects.
Associate Professor Noriyuki Kurita and Toyohashi Tech graduate students in cooperation with a researcher at the National Academy of Sciences in Ukraine, proposed new agents for protecting the cleavage site of APP from attack by γ-secretase, based on results evaluated by their state-of-the-art molecular simulations.
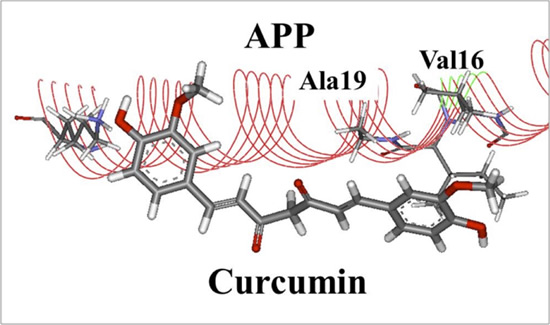
They carried out a world-first investigation into the specific interactions between a short APP peptide and curcumin derivatives, using protein-ligand docking as well as ab initio molecular simulations. Curcumin is a constituent of turmeric, which is a yellow spice commonly used in Indian cuisine. The simulated results elucidated the specific curcumin derivative and showed that it bound more strongly to APP and inhibited the binding of secretase to APP more than the other derivatives did. Moreover, some novel curcumin derivatives were proposed as potent inhibitors of Aβ peptide production. These derivatives are novel agents for suppressing AD and are proposed based on the novel concept of preventing APP cleavage. However, this does not imply that the consumption of Indian cuisine suppresses AD.
The first author, graduate student Hiromi Ishimura said, "We have also been studying the aggregation mechanism of Aβ peptides by using molecular dynamics simulations (Okamoto et al. Journal of Molecular Graphics and Modeling, 50, 113-124, 2014; Yano et al. Chemical Physics Letters, 595-596, 242-249, 2014). By combining the present ab initio molecular simulation and the molecular dynamics simulations, we will be able to elucidate the mechanism and propose potent inhibitors for suppressing the aggregation of Aβ peptides. These inhibitors are used as a therapeutic agent for ADs.”
Associate Professor Noriyuki Kurita said, “Our present molecular simulations elucidated that curcumin can bind APP and inhibit the cleavage of APP by secretase. Curcumin is a natural product contained in the root of Curcumae Rhizoma. In the laboratory of our collaborator at the National Academy of Sciences in Ukraine, our proposed curcumin derivatives will be synthesized and their effects on experimental animals with ADs will be investigated.”
Our developed state-of-the-art molecular simulations can be applied to the study of other vital proteins that play important roles in the pathogenesis of numerous diseases. In developing countries such as Ukraine, tuberculosis (TB), which is a severe infection caused by Mycobacterium tuberculosis is prevalent. In fact, about 9 million people are infected with TB worldwide and 1.5 million died from this infection in 2013. Therefore, the urgent development of novel drugs for TB is necessary. We are currently performing molecular simulations to propose novel potent inhibitors of a protein that reduces the blood level of anti-tubercular as well as anti-human immunodeficiency virus (HIV) agents. This collaborative study with Ukrainian researchers will be applied to the “Science and Technology Research Partnership for Sustainable Development Project” organized by the Japan Science and Technology Agency (JST) and the Japan International Cooperation Agency (JICA) in 2015.
Reference
Ishimura, H., Kadoya, R., Suzuki, T., Murakawa, T., Shulga, S., and Kurita, N. (2015). Specific interactions between amyloid-β peptide and curcumin derivatives: ab initio molecular simulations. Chemical Physics Letters, 633, 139-145, DOI: 10.1016/j.cplett.2015.05.023
Researcher Profile

| Name | Noriyuki Kurita |
|---|---|
| Affiliation | Department of Computer Science and Engineering |
| Title | Associate Professor |
| Fields of Research | Quantum biology / Computational science / Biological information science |
ここでコンテンツ終わりです。
